Deeply FAIR Agrigenomics: AgriSchemas, Data Integration, and the EBI GXA RDF Conversion
Deeply FAIR Agrigenomics: AgriSchemas, Data Integration, and the EBI GXA RDF Conversion
At KnetMiner, our focus is on decoding plant biology and crop traits. As a bioinformatics start-up, we build knowledge graphs to link everything from genes to phenotypes, pathways, and scientific literature. However, as biological and agrifood data continues to scale - distributed across various formats and repositories - achieving true interoperability remains a practical challenge.
This is the focus of the AgriSchemas project: an initiative led by KnetMiner and involving collaborators from the ELIXIR consortium, the FAIRAgro project, and more. AgriSchemas is designed to make agricultural, experimental, and plant data "deeply FAIR" (Findable, Accessible, Interoperable, and Reusable), where deeply means going down the dataset contents, adding to the descriptors that are about a dataset as a whole.
What is AgriSchemas?
Led by KnetMiner, AgriSchemas provides lightweight data models about agrifood and bioscience data that leverage and extend established standards like schema.org and Bioschemas. This complements highly formal approaches, such as the OBO ontologies: on the one hand, we can easily map large heterogeneous datasets into integrated semi-formal model, which would be hard to automatically map to more formal ontologies, On the other hand, we can annotate with the latter when needed (eg, to link to a precise GeneOntology function).
We use this approach to support a broad range of data types and use cases - from molecular biology and agronomic weather data to full experimental representations based on MIAPPE and ISA standards. By representing study events, experimental factors (like treatments or tillage), and observed variables (like yield or plant height) as connected nodes in a graph, AgriSchemas simplifies the integration of heterogeneous datasets into a uniform, searchable format.
The EBI Gene Expression Atlas (GXA) Challenge
A highly valuable resource for plant biologists is the European Bioinformatics Institute's Gene Expression Atlas (EBI GXA). It contains curated data detailing gene expression under different biological conditions, tissues, and treatments. However, using this tabular dataset with semantic, graph-aware tools or agents requires a structural transformation.
We needed a system that allows users to query interconnected relationships - for instance, "What happens to the expression of this specific gene in wheat when the crop faces drought stress, and what other traits does that gene map to?"
Converting GXA into RDF: Introducing agrischemas-gxapy
To solve this integration challenge, our engineering team at KnetMiner developed a dedicated Python pipeline: agrischemas-gxapy.
The agrischemas-gxapy tool is an ETL (Extract, Transform, Load) package designed to convert EBI GXA data into an RDF-based Knowledge Graph, structured cleanly around the AgriSchemas specifications.
Here is how the pipeline operates:
- Ingestion: The Python package natively extracts study descriptors, experimental designs, biological sample data, and raw gene expression metrics from GXA.
- Semantic Translation: It translates these distinct elements using AgriSchemas and Bioschemas classes. Experimental conditions become linked agri:ExperimentalFactorValue nodes, while biological inputs map directly to bioschema:Sample.
- RDF Generation: The output is a structured, linked RDF dataset. Instead of isolated spreadsheets or flat files, the GXA data becomes a connected network of relationships. Genes point to expression events, which in turn map back to explicit experimental conditions and study variables.
Seeing it in Action: Querying The Data Model with SPARQL and the Python Client
To visualize how these relationships map out practically, we have documented the core structure of the EBI GXA use case.
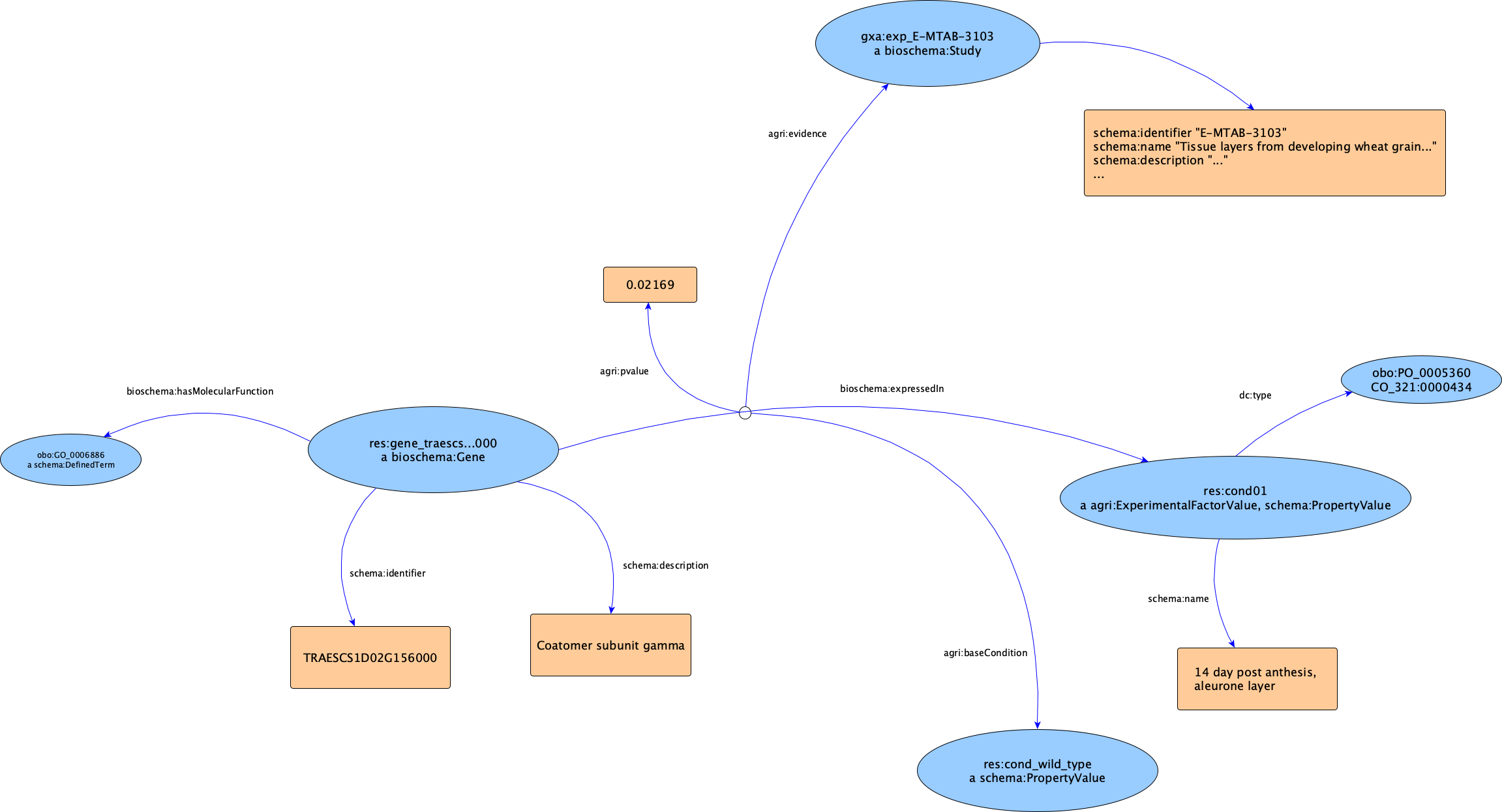
The true power of this RDF conversion is realised when querying the graph. Because the data is now fully compliant with the RDF and SPARQL semantic standards, researchers can use standard SPARQL queries to extract highly specific cross-sections of experimental data.
For example, a developer can rapidly execute a SPARQL query to retrieve all gene expression values linked to a specific environmental stress factor or disease trait across multiple studies.
You can view example queries in the data builds documentation and more details about the gene expression modelling in the GXA Use Case documentation.
If you feel intimidated by SPARQL and RDF, we have written a simple Python client library, which exposes data access functions, such as searching for studies by keywords, fetching gene expression levels with scores like p-value and fold-change.
Why This Matters for the Agritech Community
By mapping EBI’s GXA into an AgriSchemas-compliant RDF format, we enable researchers to perform semantic searches that span multiple datasets natively.
For developers, computational biologists, and bioinformaticians, this means straightforward integration of gene expression data with broader knowledge networks. You can run graph analytics to identify trait associations, trace pathways related to plant immunity, or conduct cross-variety functional annotations - all by querying a standardised knowledge graph.
Get Involved
The AgriSchemas project is open-source, and we are continually building out new FAIR dataset instances. We invite the community to explore the code, test our data builds, and view the GXA-to-RDF Python tool on our GitHub repository.
🔗 Explore the agrischemas-gxapy repository on GitHub (Note: The repository is currently hosted under the historical Rothamsted organization)
Whether you are conducting exploratory crop research, mapping candidate genes, or building the next generation of bioinformatics tools, AgriSchemas provides the semantic blueprint to bring your data together.
— The KnetMiner Team
You might also like
.png)
Introducing KnetMiner GenePage: Cross-Reference Any Gene, Powered by Graph Chat

Closing the Gap Between GWAS Peaks and Candidate Genes: Introducing Agentic KnetMiner
.png)
Deeply FAIR Agrigenomics: AgriSchemas, Data Integration, and the EBI GXA RDF Conversion
.png)
.jpg)

Graph Chat: A New Way to Explore and Understand Biological Knowledge Graphs with AI

The Future is Green: How AI in Biotech is Revolutionising Plant Genomics











